Chromatin: A Practical Approach. 1998, Oxford University Press: New York, Tokyo.
Department of Biology, Washington University, St. Louis, Missouri 63130, USA.
This chapter presents some of the techniques we have developed for the analysis of chromatin structure using a variety of nucleases to discern patterns of accessibility along the chromatin fiber. It includes mapping with DNase I, micrococcal nuclease, and restriction enzymes using embryos, larvae or adult flies as the starting material.
Genes Dev 1995 May 15;9(10):1263-1277
Department of Biology, Washington University, St. Louis, Missouri 63130, USA.
A euchromatic gene placed in the vicinity of heterochromatin by a chromosomal rearrangement generally exhibits position effect variegation (PEV), a clonally inherited pattern showing gene expression in some somatic cells but not in others. The mechanism responsible for this loss of gene expression is investigated here using fly lines carrying a P element containing the Drosophila melanogaster white and hsp26 genes. Following mobilization of the P element, a screen for variegation of white expression recovered inserts at pericentric, telomeric, and fourth chromosome regions. Previously identified suppressors of PEV suppressed white variegation of pericentric and fourth chromosome inserts but not telomeric inserts on the second and third chromosomes. This implies a difference in the mechanism for gene repression at telomeres. Heat shock-induced hsp26 expression was reduced from pericentric and fourth chromosome inserts but not from telomeric inserts. Chromatin structure analysis revealed that the variegating inserts showed a reduction in accessibility to restriction enzyme digestion in the hsp26 regulatory region in isolated nuclei. Micrococcal nuclease digests showed that pericentric inserts were packaged in a more regular nucleosome array than that observed for euchromatic inserts. These data suggest that altered chromatin packaging plays a role in PEV.
|
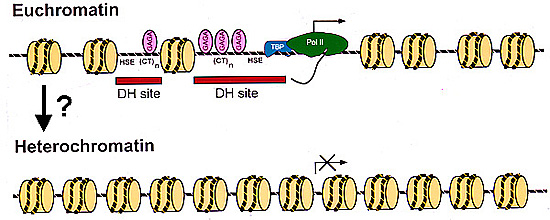
Chromatin structure of the
hsp26
transgene in euchromatin and
heterochromatin. In a euchromatic
environment, hsp26 is packaged in a specific and irregular
nucleosome array; RNA polymerases, TATA-box binding
proteins, and GAGA factors are present at their target
sequences in the open structure of the hsp26 promoter. We have
mapped this structure in some detail. |
|
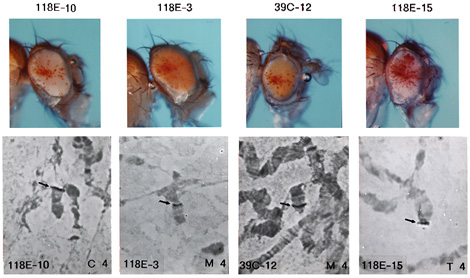
Insertion into the fourth chromosome
can result in a variegating phenotype. Mobilization of the P-element from a fully
expressing euchromatic site on the X chromosome led to
recovery of over 25 lines showing variegation which proved
to have insertion sites on the fourth chromosome.
In situ
hybridization has shown that variegating genes are inserted
not only in the pericentric and telomeric regions, but also
at many sites along the banded region of the fourth
chromosome. |
Curr Opin Genet Dev 1996 Apr;6(2):193-202
Washington University, Department of Biology, St. Louis, Missouri 63130, USA. selgin@biodec.wustl.edu
We have recently learned more about the biochemistry of heterochromatin and about how heterochromatic environments affect gene function. New findings have emphasized the distinctions between telomeric and pericentric heterochromatin in Drosophila and have suggested a mosaic structure within pericentric heterochromatin. Theories concerning the mechanism of inactivation of euchromatic genes in heterochromatic environments have been tested using transgenes inserted into heterochromatin. The current data support a competition/chromatin structure model, in which multiprotein repressor complexes compete with transcriptional activators to assemble an active or inactive chromatin structure.
J Cell Sci 1997 Sep;110( Pt 17):1999-2012
Department of Cell Biology, Yale University, New Haven, CT 06520, USA. stewart_frankel@qm.yale.edu
The actin-related proteins have been identified by virtue of their sequence similarity to actin. While their structures are thought to be closely homologous to actin, they exhibit a far greater range of functional diversity. We have localized the Drosophila actin-related protein, Arp4, to the nucleus. It is most abundant during embryogenesis but is expressed at all developmental stages. Within the nucleus Arp4 is primarily localized to the centric heterochromatin. Polytene chromosome spreads indicate it is also present at much lower levels in numerous euchromatic bands. The only other protein in Drosophila reported to be primarily localized to centric heterochromatin in polytene nuclei is heterochromatin protein 1 (HP1), which genetic evidence has linked to heterochromatin-mediated gene silencing and alterations in chromatin structure. The relationship between Arp4 and heterochromatin protein 1 (HP1) was investigated by labeling embryos and larval tissues with antibodies to Arp4 and HP1. Arp4 and HP1 exhibit almost superimposable heterochromatin localization patterns, remain associated with the heterochromatin throughout prepupal development, and exhibit similar changes in localization during the cell cycle. Polytene chromosome spreads indicate that the set of euchromatic bands labeled by each antibody overlap but are not identical. Arp4 and HP1 in parallel undergo several shifts in their nuclear localization patterns during embryogenesis, shifts that correlate with developmental changes in nuclear functions. The significance of their colocalization was further tested by examining nuclei that express mutant forms of HP1. In these nuclei the localization patterns of HP1 and Arp4 are altered in parallel fashion. The morphological, developmental and genetic data suggest that, like HP1, Arp4 may have a role in heterochromatin functions.